The Molecular Pathology Core Facility entails a full spectrum of state-of-art histopathology equipment and expertise. The facility enables investigators to study fresh, frozen, or paraffin-processed tissues and analyze sections using various histological stains and molecular probes. In addition, spatial genomics within tissue sections (GeoMx) or individual cells (CosMx) can be accomplished using multiomic platforms.
The facility is flexibly structured so that the work can be done by experienced technical staff or be trained to use the equipment themselves. Pricing has been set up accordingly.
|
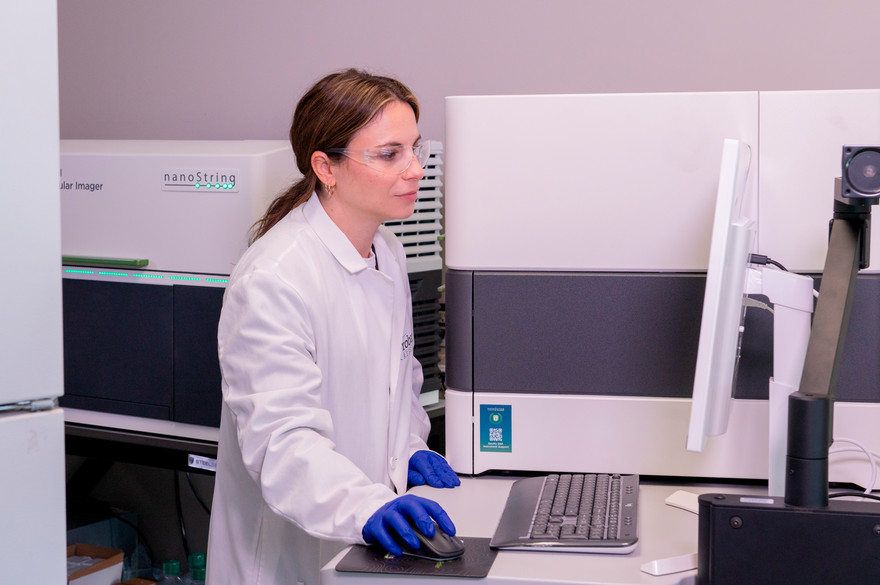
Spatial Genomics The lab has multiomic platforms for analysis of tissue sections. GeoMx Digital Spatial Profiler provides high throughput profiling of RNA and protein expression in tissue samples. CosMx is a high resolution spatial molecular imager that enables single cell spatial analysis of RNA and protein expression. |
 |
|
Immunohistrochemisty The Molecular Pathology Facility offers many staining techniques including Immunohistochemistry (IHC) or Immunofluorescence (IF), the process of localizing proteins in cells of tissue sections using the principle of antibodies binding specifically to antigens. Localization of these proteins can be customized specifically to the researchers needs. |
Laser Capture Microdissection Laser Capture Microdissection is a method for procuring pure cells from specific microscopic regions of tissue sections. Laser Capture Microdissection allows researchers to obtain homogeneous cell types and multi-cellular structures isolated from whole tissue or cytology samples. The captured cells can then be used in a wide range of downstream assays, such as gene expression analysis, genomic profiling using microarrays, or for proteomic analysis.  |
CONTACT US
Molecular PathologyRobarts Research Institute
1151 Richmond Street North
Room 4242
London, Ontario,
Canada N6A 5B7
t. 519-663-5777 ext. 24407
p. molpath@robarts.ca